Neutrons Run Enzyme’s Reactivity for Better Biofuel Production
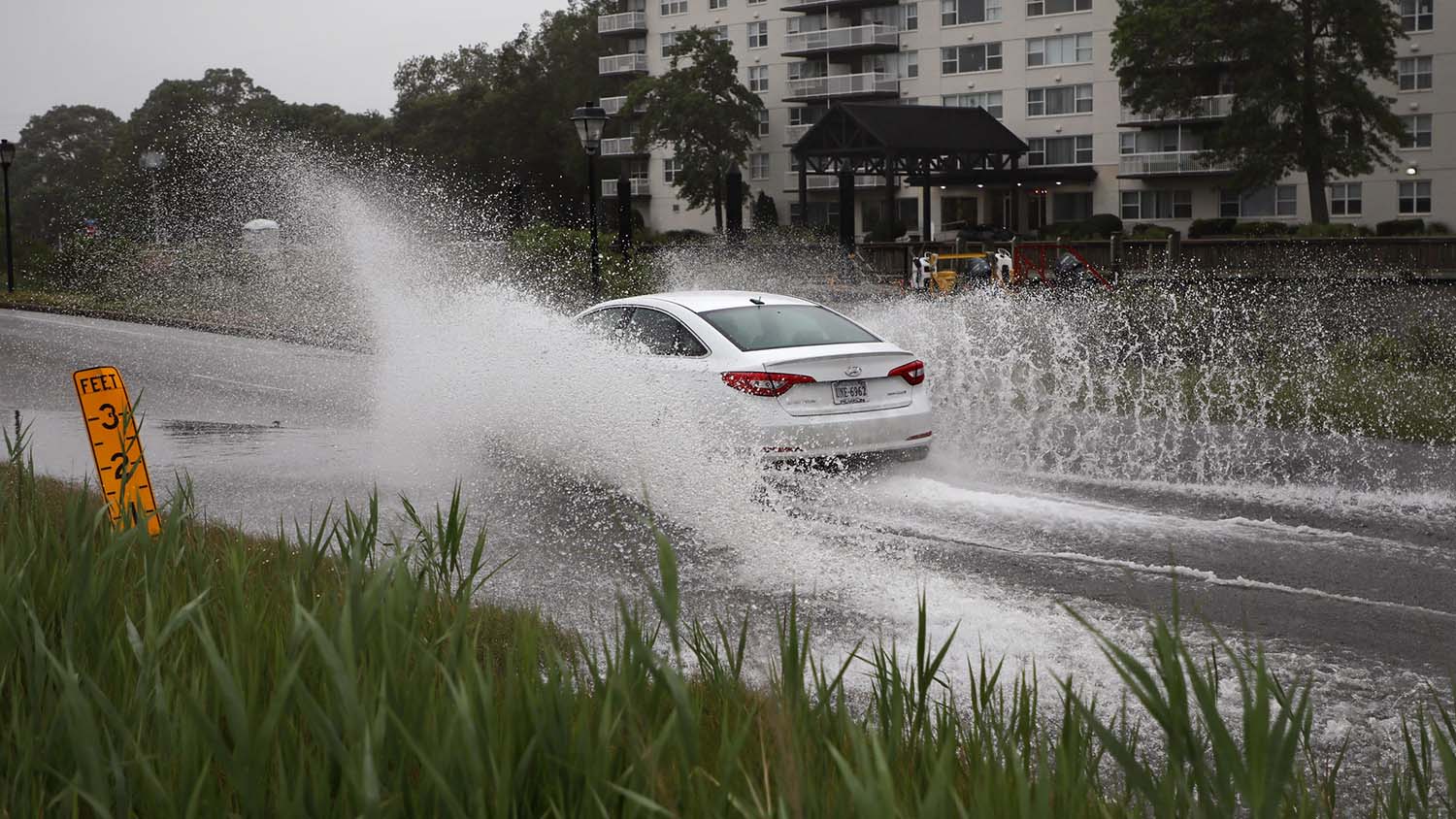
Producing biofuels like ethanol from plant materials requires various enzymes to break down the cellulosic fibers. Neutrons have identified the specifics of an enzyme-catalyzed reaction that could significantly reduce the total amount of enzymes used, improving production processes and lowering costs.
Researchers from the Department of Energy’s Oak Ridge National Laboratory and North Carolina State University used a combination of X-ray and neutron crystallography to determine the detailed atomic structure of a specialized fungal enzyme. A deeper understanding of the enzyme reactivity could also lead to improved computational models that will further guide industrial applications for cleaner forms of energy. Their results are published in the journal Angewandte Chemie International Edition.
Part of a larger family known as lytic polysaccharide monooxygenases, or LPMOs, these oxygen-dependent enzymes act in tandem with hydrolytic enzymes — which chemically break down large complex molecules with water — by oxidizing and breaking the bonds that hold cellulose chains together. The combined enzymes can digest biomass more quickly than currently used enzymes and speed up the biofuel production process.
“These enzymes are already used in industrial applications, but they’re not well understood,” said lead author Brad O’Dell, a graduate student from NC State working in the Biology and Soft Matter Division of ORNL’s Neutron Sciences Directorate. “Understanding each step in the LPMO mechanism of action will help industry use these enzymes to their full potential and, as a result, make final products cheaper.”
In an LPMO enzyme, oxygen and cellulose arrange themselves through a sequence of steps before the biomass deconstruction reaction occurs. Sort of like “on your mark, get set, go,” says O’Dell.
To better understand the enzyme’s reaction mechanism, O’Dell and coauthor Flora Meilleur, ORNL instrument scientist and an associate professor of molecular and structural biochemistry at NC State, used the IMAGINE neutron scattering diffractometer at ORNL’s High Flux Isotope Reactor to see how the enzyme and oxygen molecules were behaving in the steps leading up to the reaction—from the “resting state” to the “active state.”
The resting state, O’Dell says, is where all the critical components of the enzyme assemble to bind oxygen and carbohydrate. When electrons are delivered to the enzyme, the system moves from the resting state to the active state—i.e., from “on your mark” to “get set.”
In the active state, oxygen binds to a copper ion that initiates the reaction. Aided by X-ray and neutron diffraction, O’Dell and Meilleur identified a previously unseen oxygen molecule being stabilized by an amino acid, histidine 157.
Hydrogen is a key element of amino acids like histidine 157. Because neutrons are particularly sensitive to hydrogen atoms, the team was able to determine that histidine 157 plays a significant role in transporting oxygen molecules to the copper ion in the active site, revealing a vital detail about the first step of the LPMO catalytic reaction.
“Because neutrons allow us to see hydrogen atoms inside the enzyme, we gained essential information in deciphering the protein chemistry. Without that data, the role of histidine 157 would have remained unclear,” Meilleur said. “Neutrons were instrumental in determining how histidine 157 stabilizes oxygen to initiate the first step of the LPMO reaction mechanism.”
Their results were subsequently confirmed via quantum chemical calculations performed by coauthor Pratul Agarwal from ORNL’s Computing and Computational Sciences Directorate.
Research material preparation was supported by the ORNL Center for Structural Molecular Biology. X-ray data were collected at the Argonne National Laboratory Advanced Photon Source through access provided by the Southeast Regional Collaborative Access Team.
O’Dell says their results refine the current understanding of LPMOs for science and industry researchers.
“This is a big step forward in unraveling how LPMO’s initiate the breakdown of carbohydrates,” O’Dell said. “Now we need to characterize the enzyme’s activated state when the protein is also bound to a carbohydrate that mimics cellulose. Then we’ll have the chance to see what structural changes happen when the starting pistol is fired and the reaction takes off.”
HFIR is a DOE Office of Science User Facility. UT-Battelle manages ORNL for the Office of Science. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time.
Editor’s Note: This is a guest post submitted by Jeremy Rumsey at ORNL.
- Categories:


