Computer Modeling of WNK Kinase Inhibitors Could Give Researchers New Tools for Understanding Hypertension
For Immediate Release
Researchers from North Carolina State University have modeled and analyzed the binding modes of 210 molecules previously reported to inhibit the function of a family of enzymes involved in regulating salt and blood pressure in the human body. Their findings could help researchers better understand the complex relationships between salt regulation, hypertension and high blood pressure.
The With-No-Lysine (WNK) family of enzymes is a group of four proteins that are involved with blood pressure and body fluid regulation. These enzymes are linked with a rare and severe form of hypertension; however, their individual functions are not well understood. To better comprehend the role of each enzyme, researchers need to develop targeted molecules that can selectively shut off their function. The molecules currently in use bind with all four of the WNK proteins, preventing researchers from teasing out differences in function between each one.
Denis Fourches, assistant professor of computational chemistry, and postdoctoral researcher Melaine Kuenemann used computer modeling to characterize, analyze and visualize each of the 210 molecules known to bind with the WNK family.
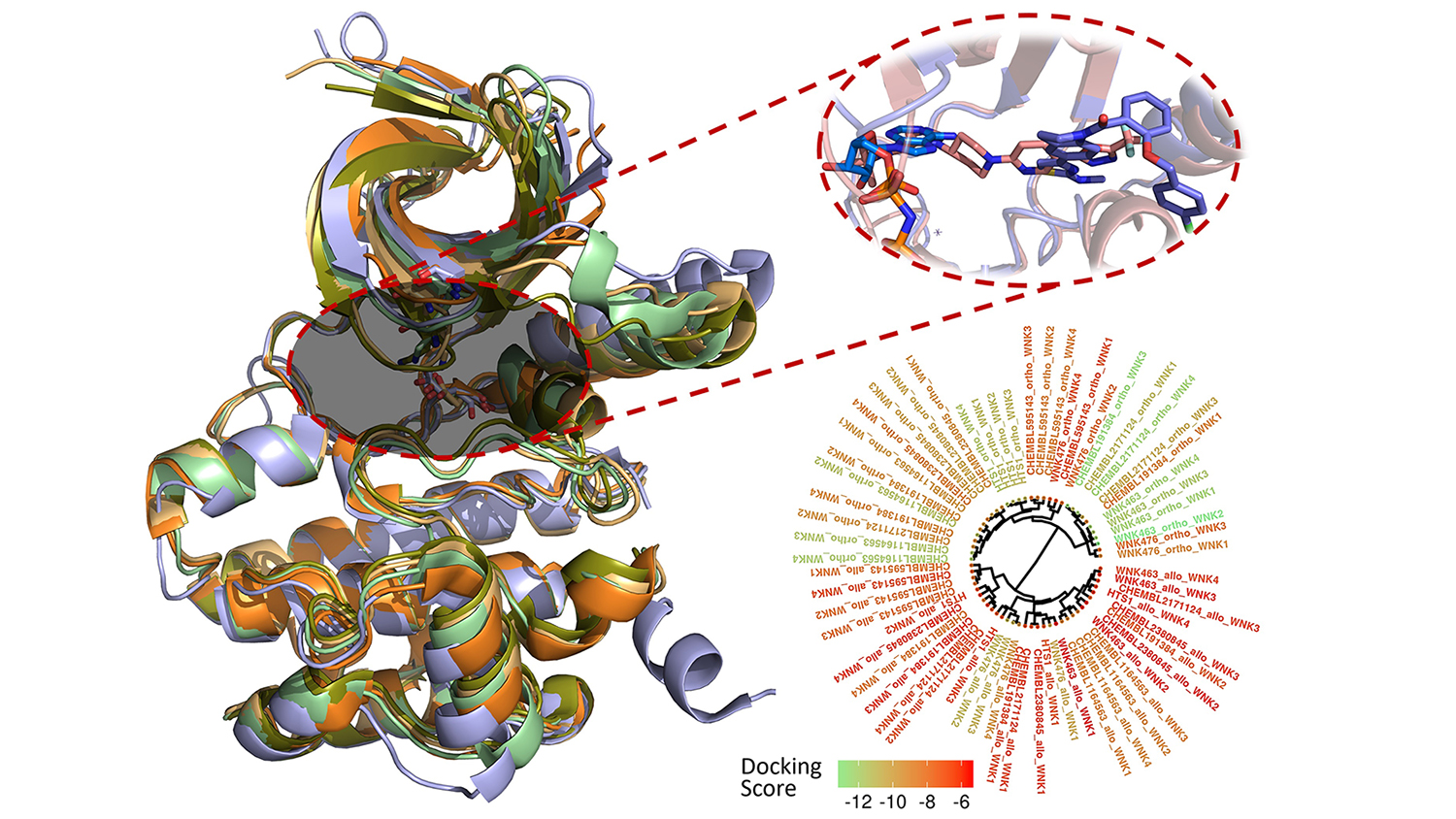
Fourches and Kuenemann created 3-D models of all 210 compounds and docked them into the binding pocket of each WNK enzyme. Then, they measured how well the compounds actually bound and analyzed their specific interactions within the four different binding pockets. They even conducted real-time molecular dynamics simulations to study how these interactions varied over time, revealing new ways to make molecules more selective toward a specific WNK enzyme.
“There aren’t that many compounds known to inhibit the WNK kinase family, and the ones we do have bind to all four enzymes,” Fourches says. “If we want to better understand the connections between these proteins and hypertension, we need to identify chemicals that can shut down one WNK kinase at a time and thus better interrogate their individual function.
“This is the first study that tries to look at the ‘big picture’ for these WNK inhibitors using state-of-the-art computer simulations. We hope that our findings will aid medicinal chemists and other researchers in designing new molecules with higher binding potency and selectivity toward each individual WNK kinase.”
The work appears in Molecular Informatics and was supported by the NC State Chancellor’s Faculty Excellence Program and the Comparative Medicine Institute at NC State.
-peake-
Note to editors: An abstract of the paper follows
“Cheminformatics Analysis of Dynamic WNK-Inhibitor Interactions”
Authors: Melaine Kuenemann, Denis Fourches, North Carolina State University
Published: Molecular Informatics
Abstract:
The With-No-Lysine (WNK) serine/threonine kinase family constitutes a unique and distinctive branch of the kinome. The four proteins of this family (WNK1/2/3/4) are involved in blood pressure regulation, body fluid, and electrolyte homeostasis. Herein, we modeled and analyzed the binding modes of all publicly-available small orthosteric and allosteric binders (including WNK463 and WNK467) experimentally tested towards any of the WNK family member. To do so, we relied on state-of-the-art cheminformatics approaches including structure-based molecular docking and molecular dynamics simulations. In particular, we computed and analyzed the (i) molecular selectivity of known inhibitors when docked in the binding site of each WNK family member, (ii) the dynamic WNK-inhibitor interactions at both orthosteric and allosteric sites to derive new structure-activity relationships, and (iii) the key specific interactions present in each binding site. This study reports on the first, cheminformatics-powered analysis of the entire chemical space of known WNK inhibitors. We discuss the conservation of critical WNK-inhibitor interactions and the existence of isoform-specific interactions that could enable the rational design of more potent and selective WNK binders.
- Categories:


