Database Helps Researchers Connect Exposures to Health Effects, Compare Diseases for Treatment
For Immediate Release
Two new studies from a group at North Carolina State University give researchers new strategies for connecting environmental exposures to human health effects.
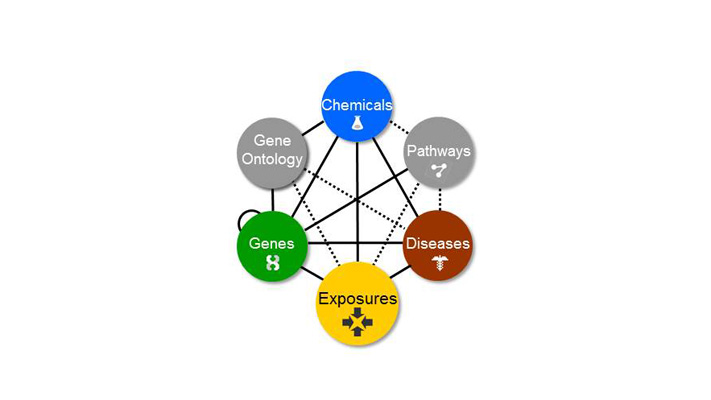
The Comparative Toxicogenomics Database (CTD) is a public database that manually curates and codes data from the scientific literature describing how environmental chemicals interact with genes to affect human health. “CTD is the only freely available database of its sort,” says Carolyn Mattingly, associate professor of biology at NC State and principal investigator of the CTD program. “It centralizes scientific data on thousands of chemicals and their relationships to genes, molecular pathways and diseases, and combines this information with tools to help scientists explore the impact of environmental exposures on human health.”
Cynthia Grondin, research scholar at NC State and lead author of the group’s study appearing today in Environmental Health Perspectives, has helped augment the database’s content to include information from exposure science articles. These data complement CTD’s experimental data with real-world exposure information on human populations and diseases.
Using an initial collection of 3,000 published articles selected for environmental exposures to humans, Ph.D.-level biocurators read the articles and hand-curated the data. They collected 54 types of data from each paper, including: the chemical involved in the exposure; demographic information on the exposed population; how the chemical was measured; and what effects were observed – including disease outcomes. Data were captured in a systematic way, incorporated into CTD and released publicly in March 2016.
“Combining this information with the more than 30 million chemical-gene-disease interactions already in CTD really expands the way users can analyze the data, by grounding experimental data in real-world contexts and providing mechanistic information to population-based studies,” says Grondin.
In another CTD study released today in PLOS ONE, Allan Peter Davis, biocuration project manager for CTD and lead author, developed a new method to find potential biological similarities between seemingly unrelated diseases. Discovering commonalities between diseases can have a big impact on drug development and treatment options for patients, as the ability to use established drugs to effectively treat several different diseases can save both time and money.
To determine whether a drug can be used to treat more than one disease, scientists look for overlaps between the set of genes that play a role in each disease: the more genes in common, the more likely the drug can be repurposed to treat both illnesses. The problem is that not all the genes involved in any one disease are always known.
Davis and the CTD team took the catalog of genes known to be associated with diseases from CTD and joined this data with a separate dataset called Gene Ontology (GO), which provides three types of descriptions for every gene: the gene product’s molecular function (what the protein does), its cellular localization (where it works in the cell) and its biological process (what roles it plays). By integrating these data sets, the CTD team produced a resource that linked over 15,000 GO annotations to 4,200 human diseases, giving them a “big picture” ability to detect biological similarities at a level above individual genes.
The team constructed a matrix that compared 4,200 human diseases and their GO annotations against one another, and then sorted the data to find the top pairs of diseases with the most significant GO overlaps. They next tested to see if they could identify drugs that could hypothetically be repurposed to treat other diseases. The group used the matrix to discover and rank 39 drugs that are currently used to treat a type of nerve cell cancer as possible therapeutics to also treat chronic B-cell leukemia. “The potential is amazing,” Davis says. “Pharmaceutical scientists can use this free resource to test new avenues for drug repositioning and potentially expanded treatment options.”
Grondin’s paper, “Advancing exposure science through chemical data curation and integration in the Comparative Toxicogenomics Database” is published in Environmental Health Perspectives. Funding was provided by NIEHS grants, ES014065, ES019604 and ES025128. Davis’ paper, “Generating Gene Ontology-Disease Inferences to Explore Mechanisms of Human Diseases at the Comparative Toxicogenomics Database” is published in PLOS ONE. This project is supported by NIEHS grants ES014065, ES023788, ES019604 and ES025128. Davis is the head of the CTD curation project. Mattingly is the director and developer of the database. NC State biologists David Reif and Jane Hoppin, research associates Thomas Wiegers, Jolene Wiegers, Daniela Sciaky, Robin J. Johnson and Benjamin King from the Mount Desert Island Biological Lab also contributed to the work.
-peake-
Note to editors: Publication information for both papers follow.
“Advancing exposure science through chemical data curation and integration in the Comparative Toxicogenomics Database”
Authors: Cynthia Grondin, Allan Peter Davis, David Reif, Jane Hoppin, Thomas Wiegers, Jolene Wiegers and Carolyn Mattingly, NC State University; Benjamin King, Mount Desert Island Biological Laboratory
Published: May 12, 2016, online in Environmental Health Perspectives
DOI: 10.1289/EHP174
“Generating Gene Ontology-Disease Inferences to Explore Mechanisms of Human Diseases at the Comparative Toxicogenomics Database”
Authors: Allan Peter Davis, Thomas Wiegers, Jolene Wiegers, Cynthia Grondin, Daniela Sciaky, Robin Johnson, and Carolyn Mattingly, North Carolina State University; and Benjamin King, The Mount Desert Island Biological Laboratory.
Published: May 12, 2016, online in PLOS ONE
- Categories:


