For Immediate Release
Researchers from North Carolina State University have discovered a way to make pinpoint changes to an enzyme-driven “assembly line” that will enable scientists to improve or change the properties of existing antibiotics as well as create designer compounds. The work is the first to efficiently manipulate which building blocks the enzyme selects in the act of synthesizing erythromycin, an important antibiotic.
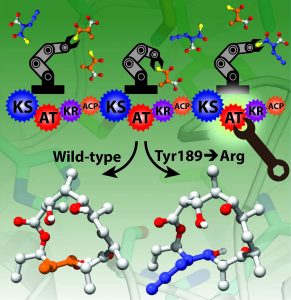
Many antibiotics are synthesized by huge sets of enzymes called polyketide synthases: a series of proteins arranged in a particular order that recruit specific small molecule building blocks to assemble the drug of interest. Picture an automobile assembly line – a car is assembled sequentially, using various interchangeable parts as it moves from one workstation to the next in line. Drug synthesis via polyketide synthases works in much the same way. Each protein module acts as a workstation responsible for selecting and adding another specific building block to the antibiotic.
Improving upon existing antibiotics is an efficient way to create new drugs in terms of both time and cost. If researchers could manipulate the function of each module in the enzyme assembly line, it would allow them to design man-made molecules, thus fine-tuning the pharmacological properties of a drug. However, no one had been able to discover how to make small tweaks to an enzyme module in order to completely change which building blocks are selected during the assembly process.
Gavin Williams, associate professor of bio-organic chemistry at NC State and corresponding author of a paper describing the work, along with former Ph.D. student and first author Irina Koryakina, former Ph.D student John McArthur and current Ph.D. student Christian Kasey, set out to discover how to make an enzyme module in erythromycin select a man-made building block.
The group looked at a protein module designated Ery6, which is the sixth module, or workstation on the assembly line, for erythromycin. They found that the module identifies and installs a methyl group building block. By genetically altering the active site of the enzyme they were able to make changes to single amino acids in that area, engineering the enzyme to reject the naturally occurring substrate, or building block, that it normally selected in favor of a man-made substrate the researchers preferred.
“We have engineered the incorporation of a non-natural substrate by changing the building block specificity of Ery6,” Kasey says. “Because we have completely changed the specificity only at the sixth module, we know that the chemistry at a defined part of the molecule will be changed. So if we move to a scenario where these compounds are generated in environments that may have both the natural and man-made substrates available to them, we know that only module six will select the substrate we want.”
“Previously, most changes to these compounds have been in the form of swapping out an entire enzyme module, rather than tweaking functionality within it,” Williams says. “Instead of a hatchet, our method is more surgical, making small but impactful changes to the module that won’t change its overall function while allowing us to fine tune the portions of the compound that we select.
“We want to apply this same approach to alter other groups in the structure so that we could diversify and modify other properties of the antibiotic. We believe that this approach will prove a powerful tool in constructing new designer compounds with pinpoint accuracy.”
The work appears in ACS Chemical Biology and was funded in part by the National Science Foundation (CHE-1151299) and the National Institutes of Health (GM104258). Koryakina is currently at Intrexon Corporation, and McArthur is currently at the University of California, Davis. Joseph Chemler, Shasha Li, Douglas Hansen and David Sherman from the University of Michigan, Ann Arbor also contributed to the work.
-peake-
Note to editors: An abstract of the paper follows.
“Inversion of extender unit selectivity in the erythromycin polyketide synthase by acyltransferase domain engineering”
DOI: 10.1021/acschembio.6b00732
Authors: Irina Koryakina, NC State and Intrexon Corp.; John McArthur, NC State and UC-Davis; Christian Kasey and Gavin Williams, NC State University; Andrew Lowell, Joseph Chemler, Shasha Li, Douglas Hansen and David Sherman, University of Michigan Ann Arbor
Published: ACS Chemical Biology
Abstract:
Acyltransferase (AT) domains of polyketide synthases (PKSs) select extender units for incorporation into polyketides and dictate large portions of the structures of clinically relevant natural products. Accordingly, there is significant interest in engineering the substrate specificity of PKS ATs in order to site-selectively manipulate polyketide structure. However, previous attempts to engineer ATs have yielded mutant PKSs with relaxed extender unit specificity, rather than an inversion of selectivity from one substrate to another. Here, by directly screening the extender unit selectivity of mutants from active site saturation libraries of an AT from the prototypical PKS, 6- deoxyerythronolide B synthase, a set of single amino acid substitutions was discovered that dramatically impact the selectivity of the PKS with only modest reductions of product particular substitution (Tyr189Arg) inverted the selectivity of the wild-type PKS from its natural substrate towards a non-natural alkynyl-modified extender unit while maintaining more than twice the activity of the wild-type PKS with its natural substrate. The strategy and mutations described herein form a platform for combinatorial biosynthesis of site-selectively modified polyketide analogues that are modified with nonnatural and non-native chemical functionality.
- Categories:



