Researchers Use DNA ‘Clews’ to Shuttle CRISPR-Cas9 Gene-Editing Tool into Cells
For Immediate Release
Researchers from North Carolina State University and the University of North Carolina at Chapel Hill have for the first time created and used a nanoscale vehicle made of DNA to deliver a CRISPR-Cas9 gene-editing tool into cells in both cell culture and an animal model.
 The CRISPR-Cas system, which is found in bacteria and archaea, protects bacteria from invaders such as viruses. It does this by creating small strands of RNA called CRISPR RNAs, which match DNA sequences specific to a given invader. When those CRISPR RNAs find a match, they unleash Cas9 proteins that cut the DNA. In recent years, the CRISPR-Cas system has garnered a great deal of attention in the research community for its potential use as a gene editing tool – with the CRISPR RNA identifying the targeted portion of the relevant DNA, and the Cas protein cleaving it.
The CRISPR-Cas system, which is found in bacteria and archaea, protects bacteria from invaders such as viruses. It does this by creating small strands of RNA called CRISPR RNAs, which match DNA sequences specific to a given invader. When those CRISPR RNAs find a match, they unleash Cas9 proteins that cut the DNA. In recent years, the CRISPR-Cas system has garnered a great deal of attention in the research community for its potential use as a gene editing tool – with the CRISPR RNA identifying the targeted portion of the relevant DNA, and the Cas protein cleaving it.
But for Cas9 to do its work, it must first find its way into the cell. This work focused on demonstrating the potential of a new vehicle for directly introducing the CRISPR-Cas9 complex – the entire gene-editing tool – into a cell.
“Traditionally, researchers deliver DNA into a targeted cell to make the CRISPR RNA and Cas9 inside the cell itself – but that limits control over its dosage,” says Chase Beisel, co-senior author of the paper and an assistant professor in the department of chemical and biomolecular engineering at NC State. “By directly delivering the Cas9 protein itself, instead of turning the cell into a Cas9 factory, we can ensure that the cell receives the active editing system and can reduce problems with unintended editing.”
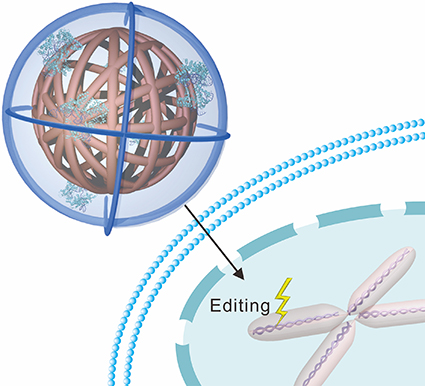
“Our delivery mechanism resembles a ball of yarn, or clew, so we call it a nanoclew,” says Zhen Gu, co-senior author of the paper and an assistant professor in the joint biomedical engineering program at NC State and UNC-CH. “Because the nanoclew is made of a DNA-based material, it is highly biocompatible. It also self-assembles, which makes it easy to customize.”
The nanoclews are made of a single, tightly-wound strand of DNA. The DNA is engineered to partially complement the relevant CRISPR RNA it will carry, allowing the CRISPR-Cas9 complex – a CRISPR RNA bound to a Cas9 protein — to loosely attach itself to the nanoclew. “Multiple CRISPR-Cas complexes can be attached to a single nanoclew,” says Wujin Sun, lead author of the study and Ph.D. student in Gu’s lab.
When the nanoclew comes into contact with a cell, the cell absorbs the nanoclew completely – swallowing it and wrapping it in a protective sheath called an endosome. But the nanoclews are coated with a positively-charged polymer that breaks down the endosome, setting the nanoclew free inside the cell. The CRISPR-Cas9 complexes can then free themselves from the nanoclew to make their way to the nucleus. And once a CRISPR-Cas9 complex reaches the nucleus, gene editing begins.
To test the nanoclew CRISPR-Cas delivery system, the researchers treated cancer cell cultures and tumors in mice. The relevant cancer cells had been modified to express a fluorescent protein. In short, they glowed. The CRISPR RNAs on the nanoclews were designed to target the DNA in the cancer cell that was responsible for making the fluorescent proteins. If the glowing stopped, the nanoclews worked.
“And they did work. More than a third of cancer cells stopped expressing the fluorescent protein,” Beisel says.
“This study is a proof of concept, and additional work needs to be done – but it’s very promising,” Gu says.
The paper, “Self-Assembled DNA Nanoclews for the Efficient Delivery of CRISPR–Cas9 for Genome Editing,” is published in the journal Angewandte Chemie. Co-authors include Jordan Hall, an undergraduate at NC State; and Wenyan Ji, Quanyin Hu and Chao Wang of the joint biomedical engineering program. The work was supported by the NC TraCS, NIH’s Clinical and Translational Science Awards at UNC-CH, grant number 1UL1TR001111 and by National Science Foundation under grant MCB-1452902.
-shipman-
Note to Editors: The study abstract follows.
“Self-Assembled DNA Nanoclews for the Efficient Delivery of CRISPR–Cas9 for Genome Editing”
Authors: Wujin Sun, Wenyan Ji, Quanyin Hu, Chao Wang and Zhen Gu, North Carolina State University and the University of North Carolina at Chapel Hill; Jordan M. Hall and Chase L. Beisel, North Carolina State University
Published: Aug. 27, Angewandte Chemie
DOI: 10.1002/anie.201506030
Abstract: CRISPR–Cas9 represents a promising platform for genome editing, yet means for its safe and efficient delivery remain to be fully realized. A novel vehicle that simultaneously delivers the Cas9 protein and single guide RNA (sgRNA) is based on DNA nanoclews, yarn-like DNA nanoparticles that are synthesized by rolling circle amplification. The biologically inspired vehicles were efficiently loaded with Cas9/sgRNA complexes and delivered the complexes to the nuclei of human cells, thus enabling targeted gene disruption while maintaining cell viability. Editing was most efficient when the DNA nanoclew sequence and the sgRNA guide sequence were partially complementary, offering a design rule for enhancing delivery. Overall, this strategy provides a versatile method that could be adapted for delivering other DNA-binding proteins or functional nucleic acids.
- Categories:


